
PDBMine - Data
Computation and Visualization for Protein Analysis
Why Use PDBMine - Data?
The cause of any disease can be traced back to misfunction, hyperfunction, and dysfunction of one or more proteins at the cellular level. A structural understanding of proteins leads to a better understanding of its biological functions, necessary to develop effective medicine or vaccine. Our computational approach is one of the most cost-effective and time-efficient compared to traditional methods.
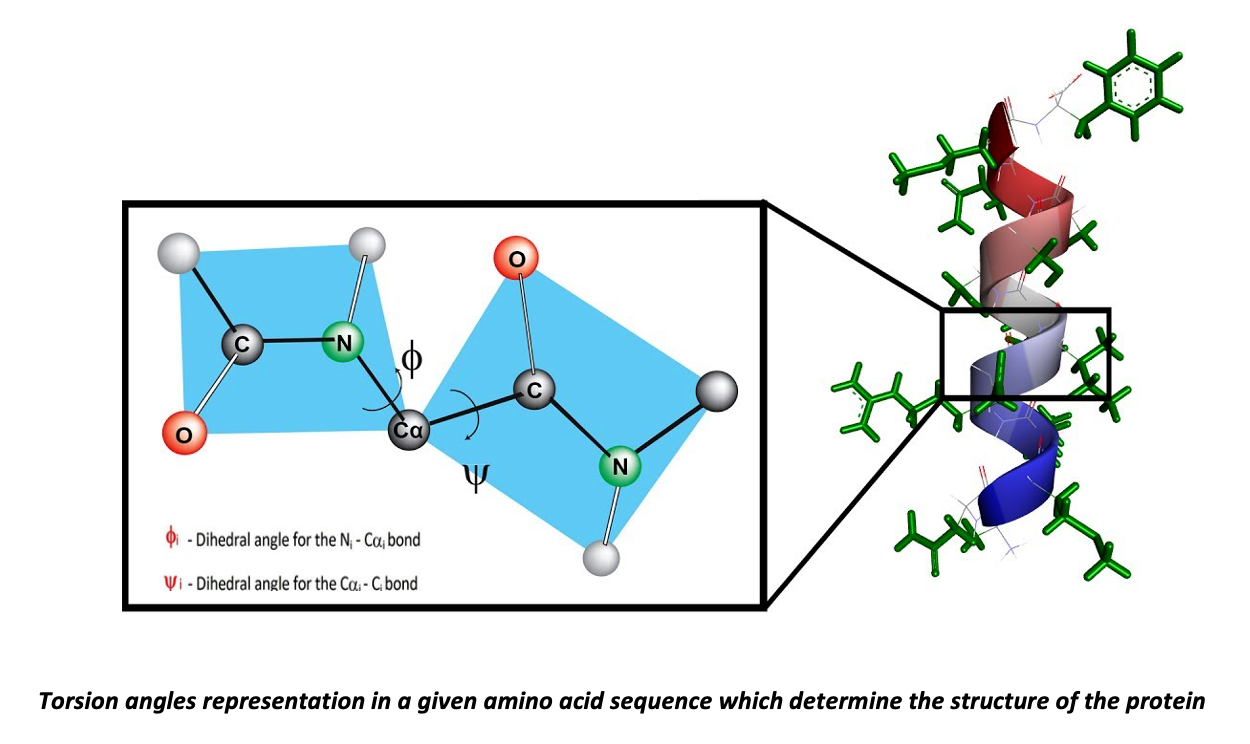
PDBMine discovers existing protein structures with a similar k-mer sequence of the protein; this structural similarity helps construct its fragmented structure. PDBMine returns torsion angles, which our website clusters and displays in a scatter plot. Furthermore, PDBMine - Data can use these angles to generate a pdb file modeling the potential structure of the protein.
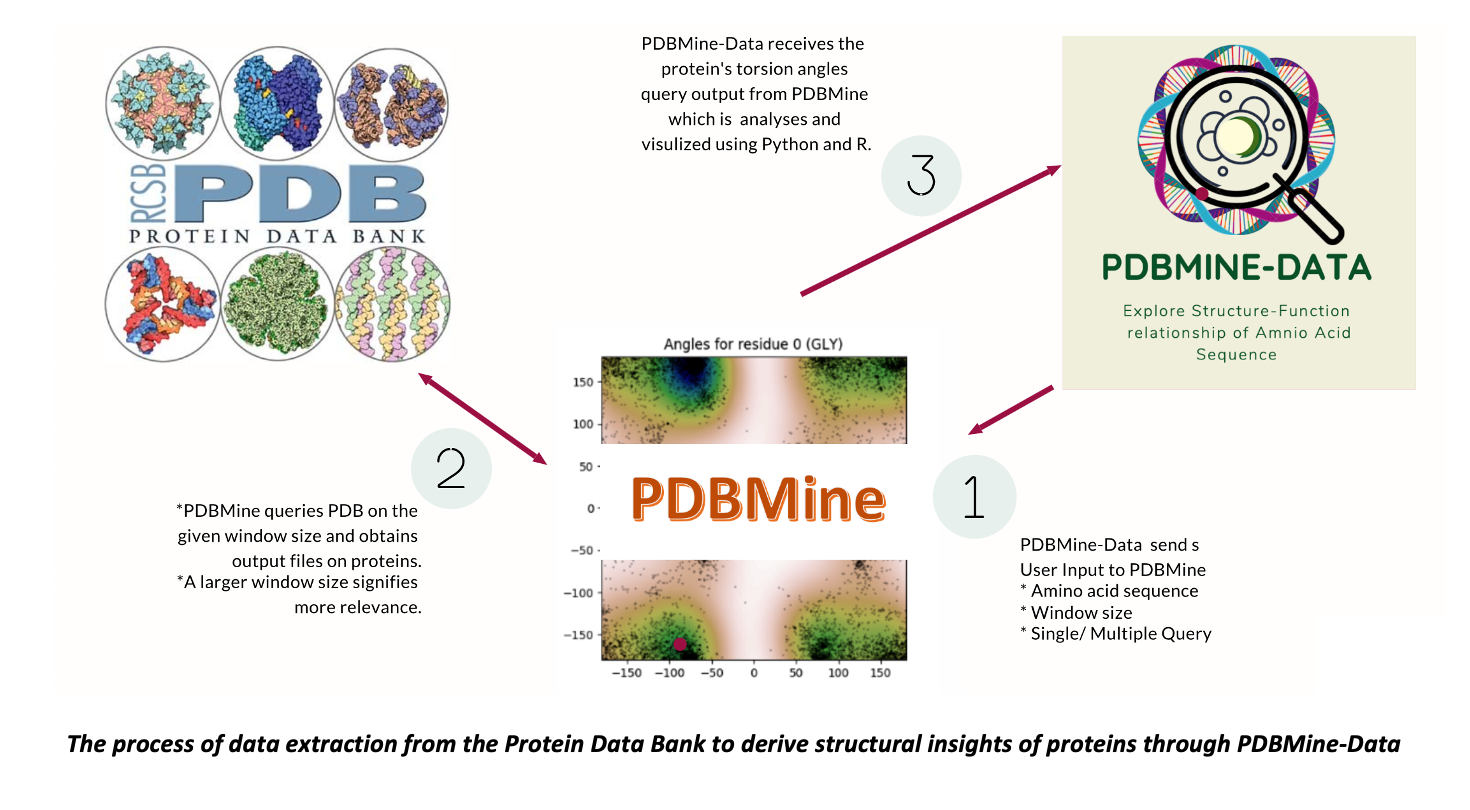
How to Use PDBMine - Data
Our website allows users to go from a sequence to a possible structure in 3 easy steps! First, create a new project with the desired protein sequence and window size.
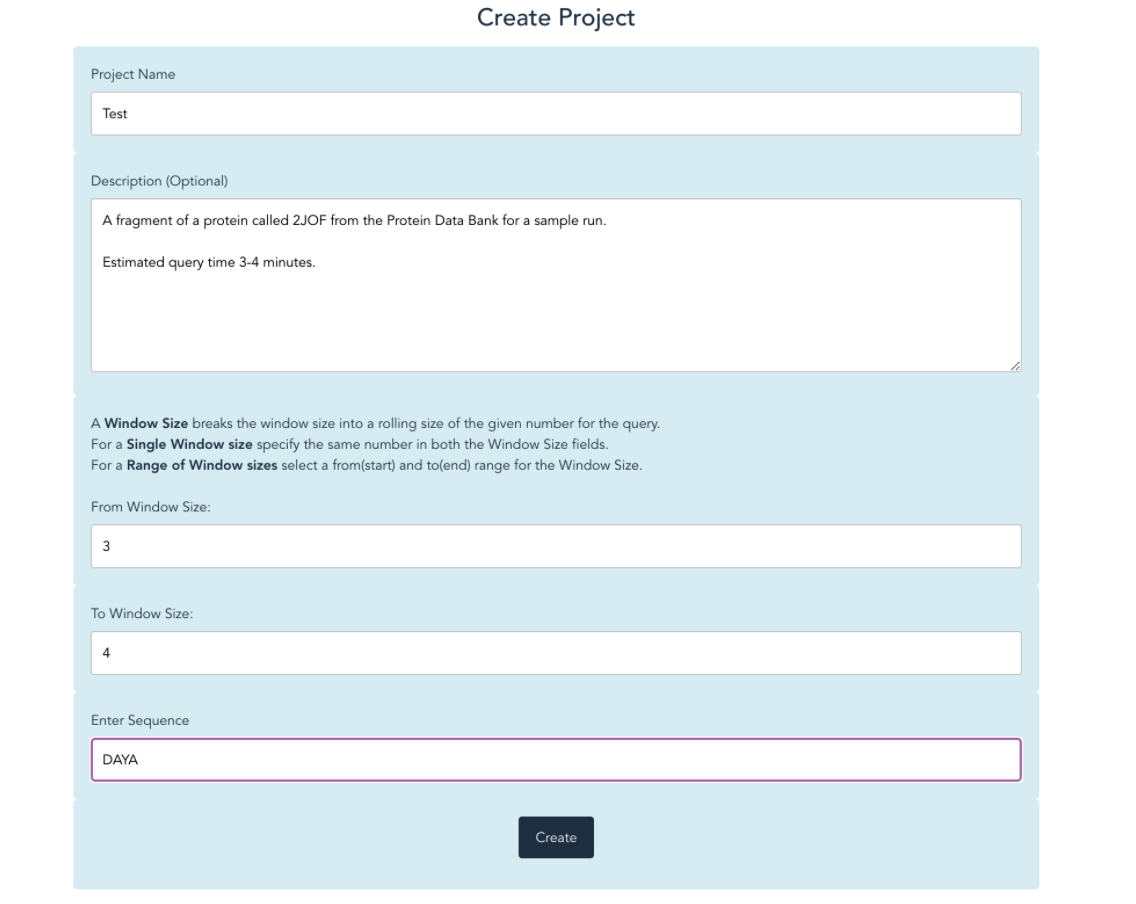
Second, find the new project in your Projects list and view the results. When the results are finished processing, you will be able to view scatter plots of the phi and psi angles of each residue in the sequence.
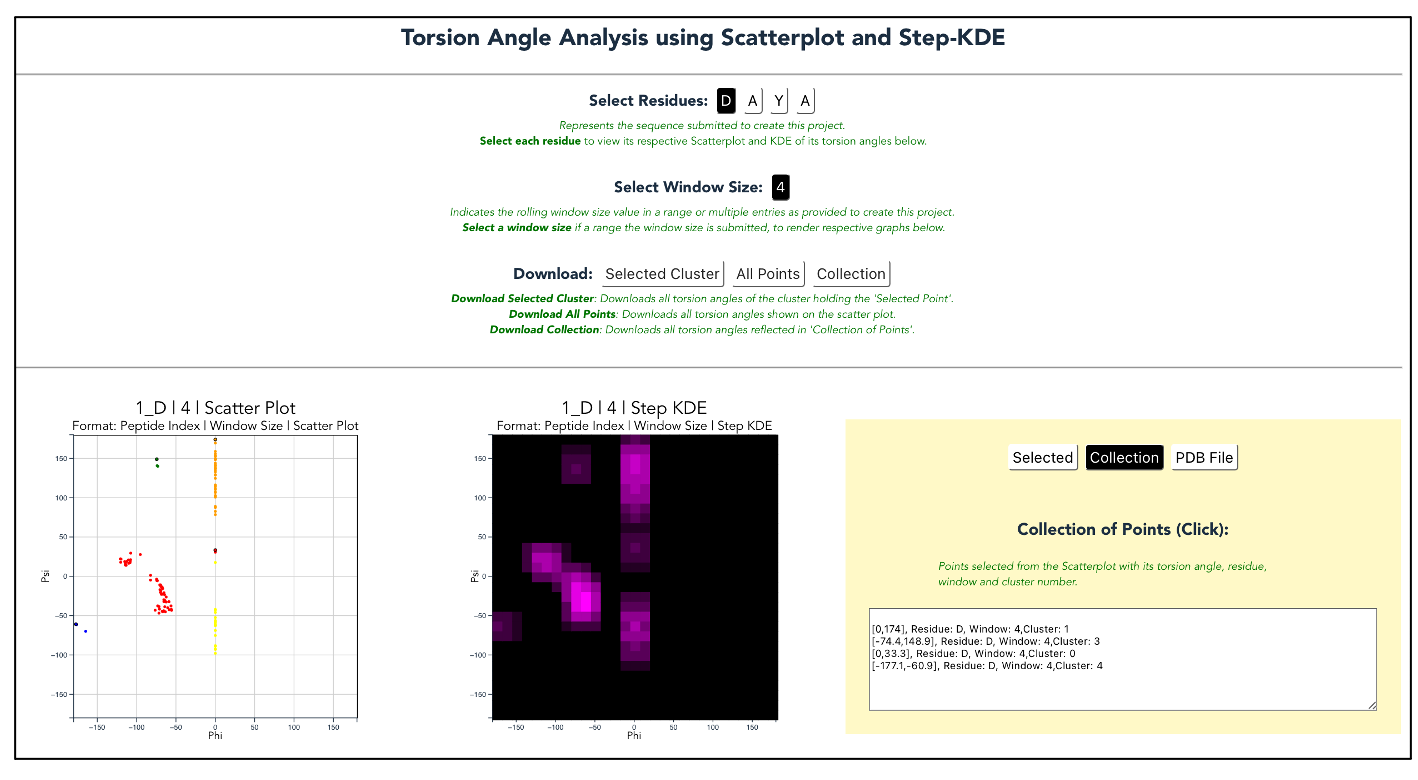
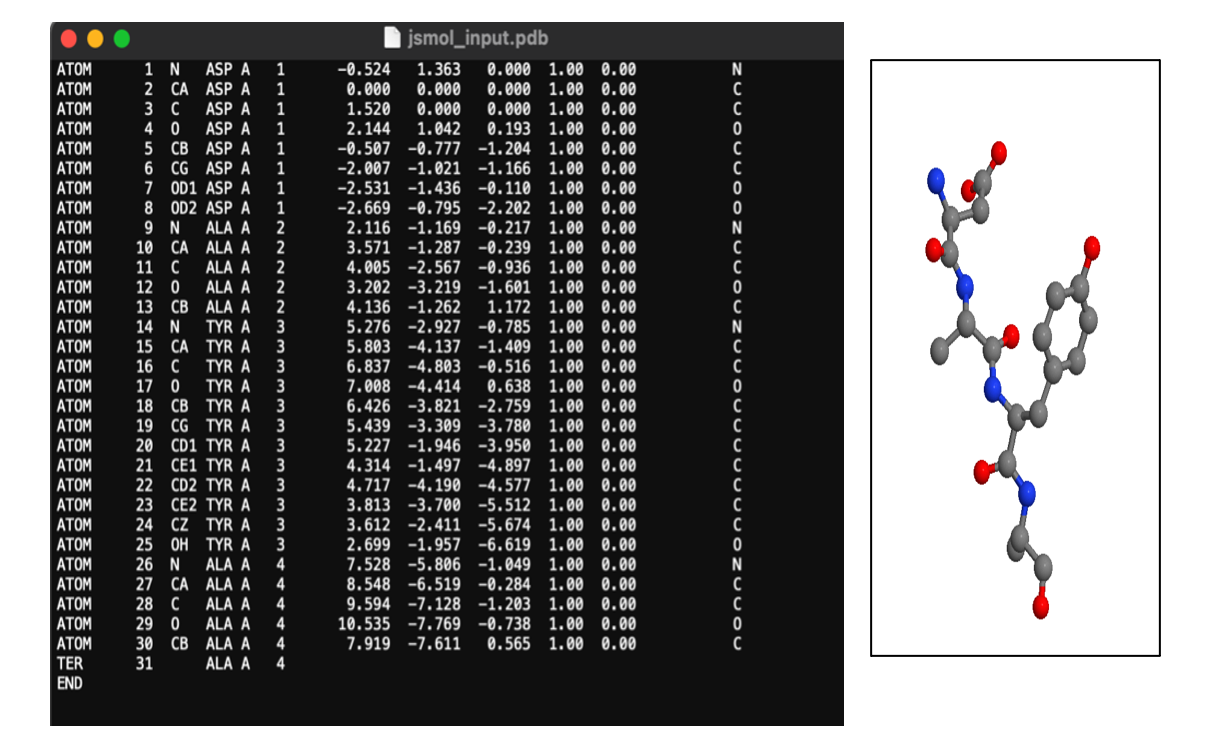
Third, select the set of angles you wish to use and click “generate.” This will open download a pdb file of your chosen structure that you can view in your favorite pdb modeling program.
About the team

Learn More
If you want to explore PDBMine - Data in more detail, our github repo can be found here